Takes as input a sefraModel object following a call to sampler and plots traces for desired parameters.
Arguments
- object
sefraModelobject.- pars
character vector of posterior parameter samples to be extracted.
- labels
character or expression vector of labels
- ...
(not used)
Examples
library(cmdstanr)
library(ggplot2)
#> Warning: package 'ggplot2' was built under R version 4.5.2
CODE <- "data{ int n; vector[n] x; }
parameters{ real mu; }
model{ x ~ normal(mu, 1.0);}
generated quantities{ vector[n] x_sim; real x_sim_sum;
for (i in 1:n) x_sim[i] = normal_rng(mu, 1.0); x_sim_sum = sum(x_sim);}\n"
ff <- write_stan_file(CODE)
mdl <- cmdstan_model(stan_file = ff); rm(ff)
n <- 20
x <- rnorm(n, 0, 2)
mdl_fit <- mdl$sample(data = list(n = n, x = x), init = function() list(mu = 0), chains = 1)
#> Running MCMC with 1 chain...
#>
#> Chain 1 Iteration: 1 / 2000 [ 0%] (Warmup)
#> Chain 1 Iteration: 100 / 2000 [ 5%] (Warmup)
#> Chain 1 Iteration: 200 / 2000 [ 10%] (Warmup)
#> Chain 1 Iteration: 300 / 2000 [ 15%] (Warmup)
#> Chain 1 Iteration: 400 / 2000 [ 20%] (Warmup)
#> Chain 1 Iteration: 500 / 2000 [ 25%] (Warmup)
#> Chain 1 Iteration: 600 / 2000 [ 30%] (Warmup)
#> Chain 1 Iteration: 700 / 2000 [ 35%] (Warmup)
#> Chain 1 Iteration: 800 / 2000 [ 40%] (Warmup)
#> Chain 1 Iteration: 900 / 2000 [ 45%] (Warmup)
#> Chain 1 Iteration: 1000 / 2000 [ 50%] (Warmup)
#> Chain 1 Iteration: 1001 / 2000 [ 50%] (Sampling)
#> Chain 1 Iteration: 1100 / 2000 [ 55%] (Sampling)
#> Chain 1 Iteration: 1200 / 2000 [ 60%] (Sampling)
#> Chain 1 Iteration: 1300 / 2000 [ 65%] (Sampling)
#> Chain 1 Iteration: 1400 / 2000 [ 70%] (Sampling)
#> Chain 1 Iteration: 1500 / 2000 [ 75%] (Sampling)
#> Chain 1 Iteration: 1600 / 2000 [ 80%] (Sampling)
#> Chain 1 Iteration: 1700 / 2000 [ 85%] (Sampling)
#> Chain 1 Iteration: 1800 / 2000 [ 90%] (Sampling)
#> Chain 1 Iteration: 1900 / 2000 [ 95%] (Sampling)
#> Chain 1 Iteration: 2000 / 2000 [100%] (Sampling)
#> Chain 1 finished in 0.1 seconds.
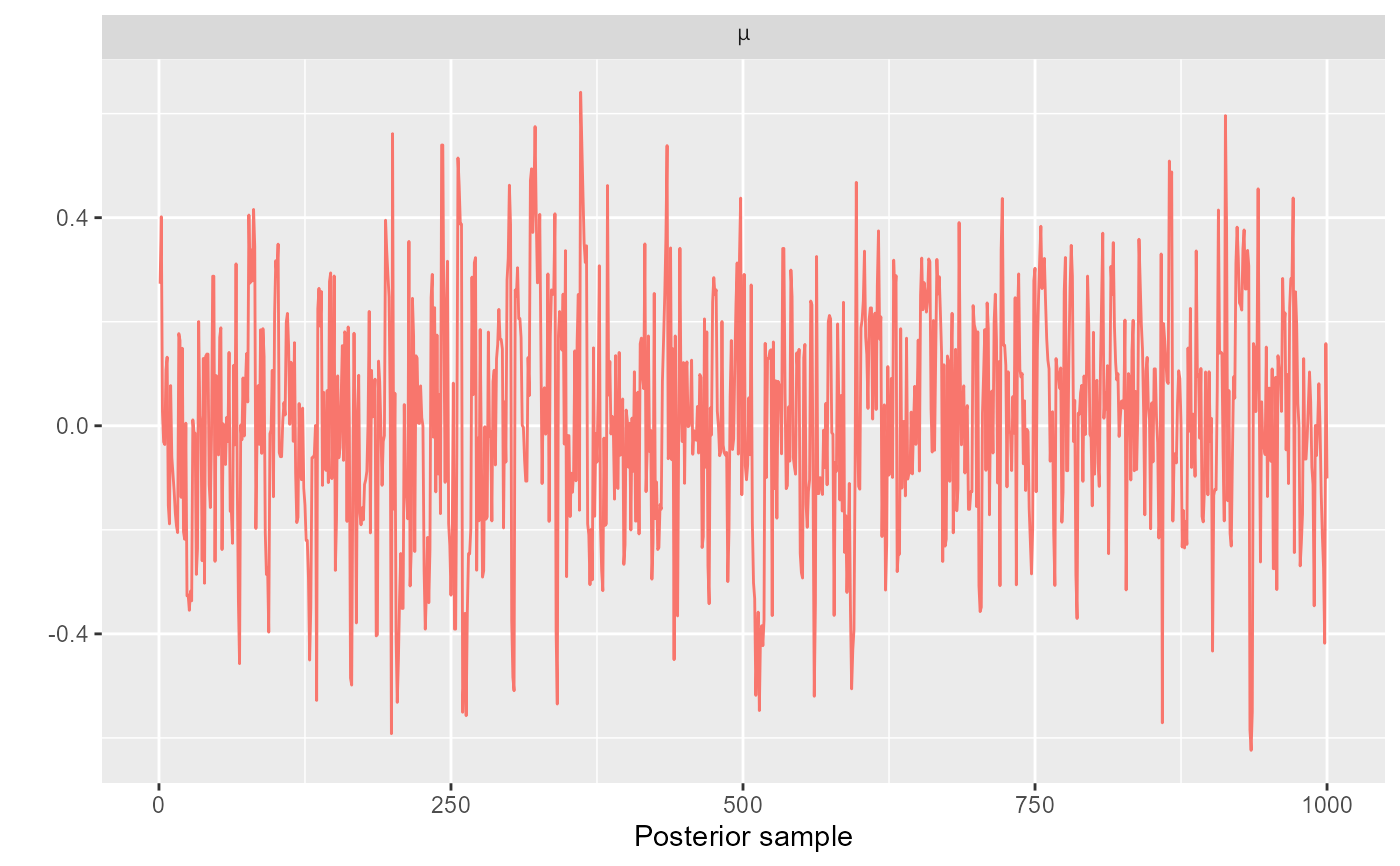
trace_plot(mdl_fit, pars = "mu", labels = list(1)) + facet_wrap(~"mu", labeller = label_parsed)